The Structural Biology, Biochemistry and Biophysics Program
The Structural Biology, Biochemistry and Biophysics Program (SB3) at the University of Connecticut is one of four areas of concentration within the Molecular and Cell Biology Field of Study, offering both PhD and MS degrees. We study the structure, function, and interactions of biological macromolecules.
Research Focus
Areas of research focus on the structure, function, and interactions of biological macromolecules. Faculty expertise includes structural biology (cryoelectron microscopy, NMR, and x-ray crystallography), computational biology, and advanced biochemical and biophysical techniques. Experimental systems in SB3 range from macromolecules and macromolecular complexes to organellar and cellular models.
WHO WE ARE
CONTACT
Eric May, Program Head
Phone: (860) 486-0484
Email: eric.may@uconn.edu
SB3 NEWS
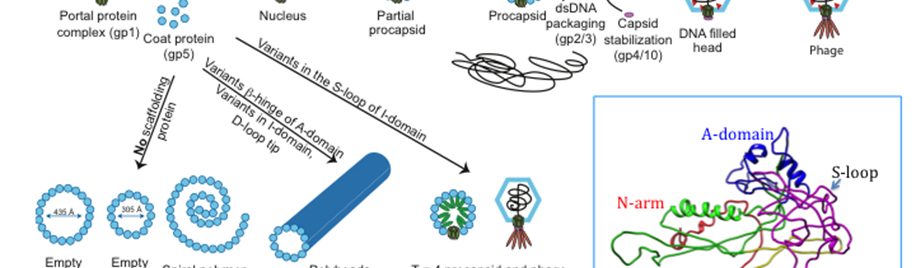
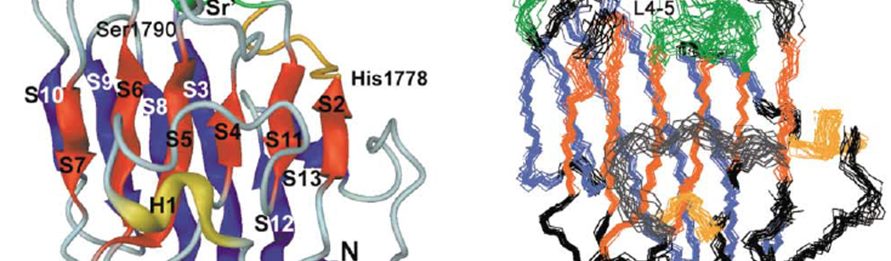
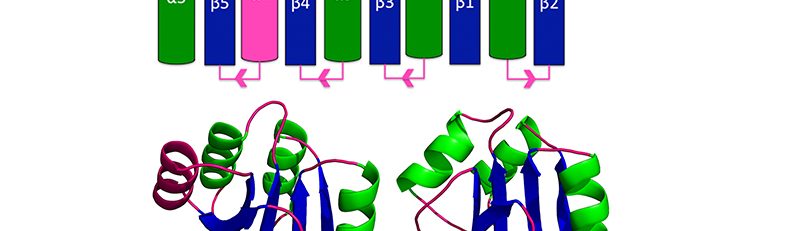
- Alexandrescu/Teschke Labs PublicationTemplated trimerization of the phage L decoration protein on capsids Protein Sci.Posted on March 19, 2025
- MCB/SB3 Welcome New FacultyMCB is excited to introduce our two newest faculty members, Dylan Murray and Kristin Ramsey Dylan Murray joins the Department of Molecular and Cell Biology as an assistant professor. Murray earned his doctorate degree in molecular biophysics from Florida State University and worked as a postdoctoral fellow at the National Institutes of Health. Murray’s research interest […]Posted on September 3, 2024
- Alder Awarded Reinhard-Frank Foundation GrantNathan Alder, along with collaborator Doron Rapaport from the University of Tübingen (Germany), has received an award from the Reinhard-Frank Foundation for research on mitochondria-targeted bioactive compounds. Support from this foundation is designed to advance novel research that builds upon existing research strengths and promotes sustained partnership between participating institutions. The supported research will explore […]Posted on February 13, 2023